Cophenetic metrics for phylogenetic trees, after Sokal and Rohlf
Correlation between metrics over phylogenetic trees with 10 leaves (Simulation)
Non-necessary binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 | 1.0 | 0.976490710347 | 0.779302323399 | 0.821463184973 | 0.343396219683 |
| Coph p=2 | | 1.0 | 0.811423373172 | 0.862097549527 | 0.361572343727 |
| Nsp p=1 | | | 1.0 | 0.971139418548 | 0.504290713833 |
| Nsp p=2 | | | | 1.0 | 0.465931915873 |
| RF | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 |  |  |  |  |
| Coph p=2 | |  |  |  |
| Nsp p=1 | | | 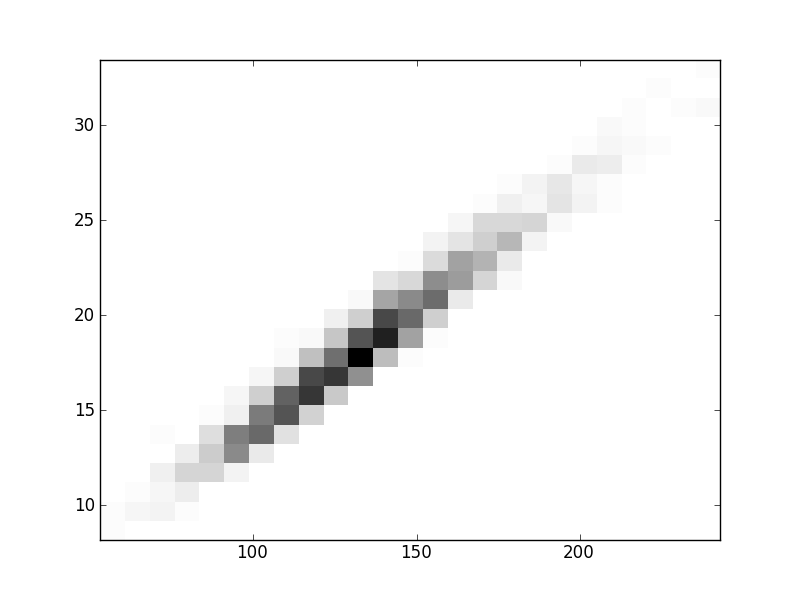 |  |
| Nsp p=2 | | | |  |
Binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 | 1.0 | 0.975610661412 | 0.870082165649 | 0.887771828039 | 0.140522058863 | 0.152808674647 | 0.260271979703 |
| Coph p=2 | | 1.0 | 0.893731005636 | 0.918727563761 | 0.179329558323 | 0.198146286331 | 0.262087574887 |
| Nsp p=1 | | | 1.0 | 0.972779078961 | 0.292995937421 | 0.298535964906 | 0.326602411291 |
| Nsp p=2 | | | | 1.0 | 0.288220797993 | 0.30756409948 | 0.296825481517 |
| Nunsp p=1 | | | | | 1.0 | 0.958927929611 | 0.336375774237 |
| Nunsp p=2 | | | | | | 1.0 | 0.349566705325 |
| RF | | | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 | 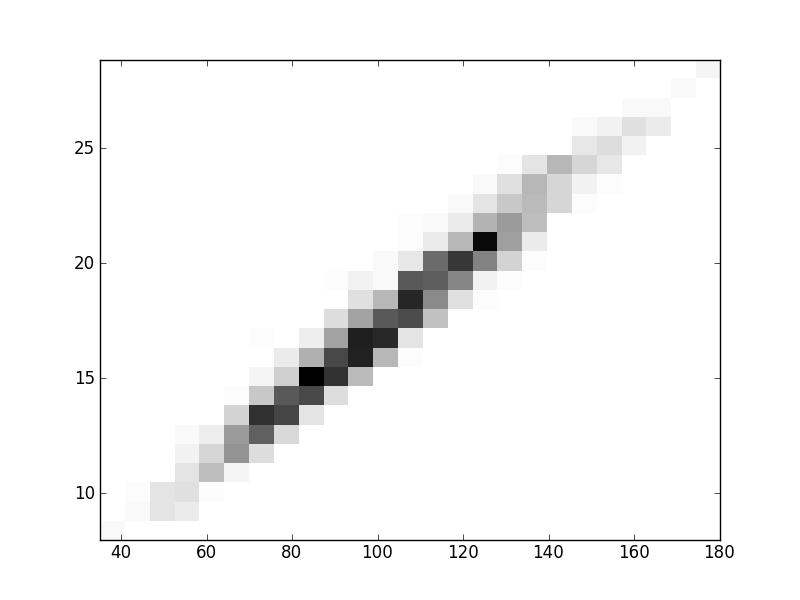 |  |  | 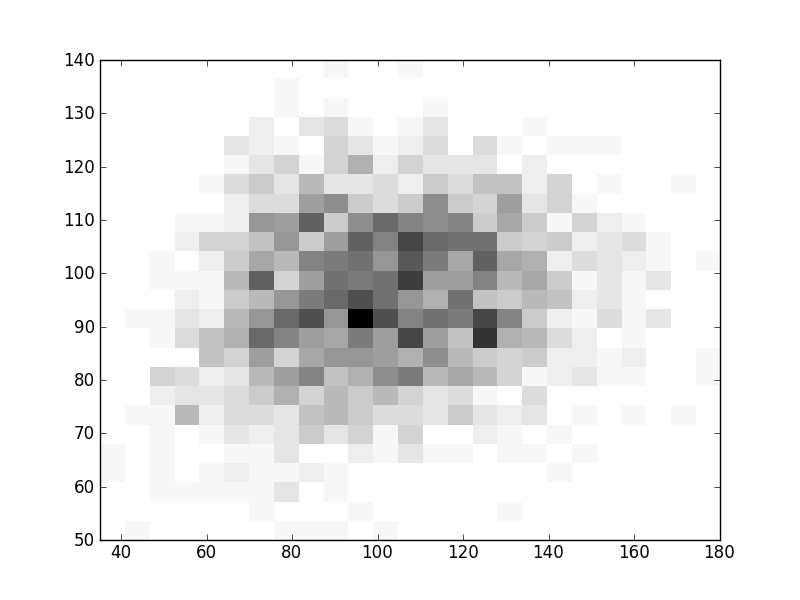 |  |  |
| Coph p=2 | |  |  | 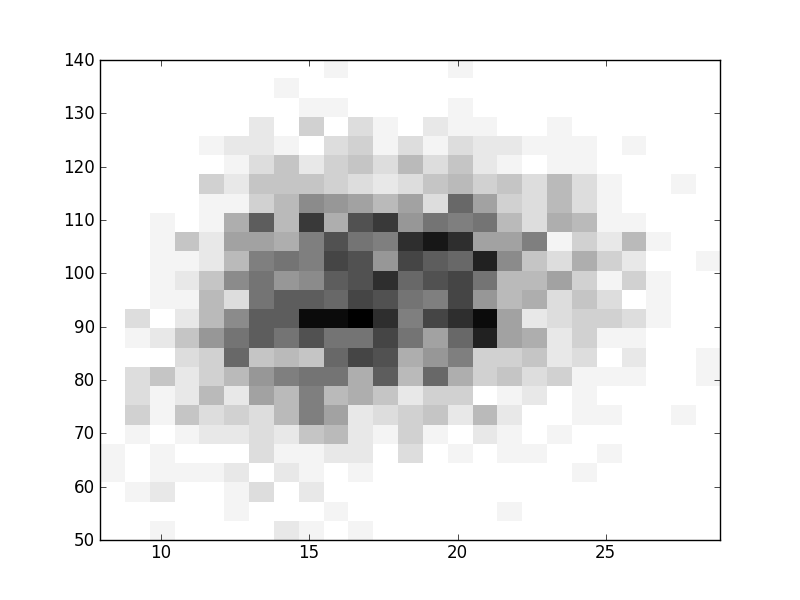 |  |  |
| Nsp p=1 | | |  |  |  | 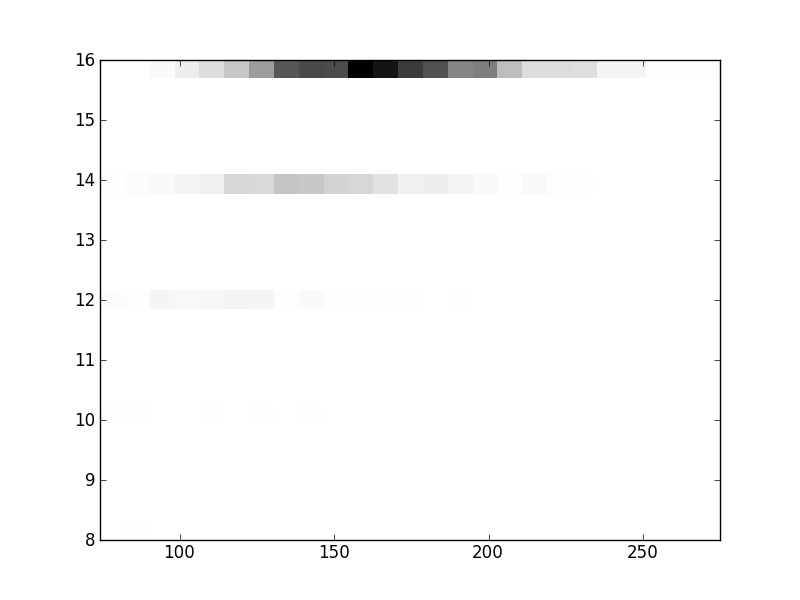 |
| Nsp p=2 | | | |  |  | 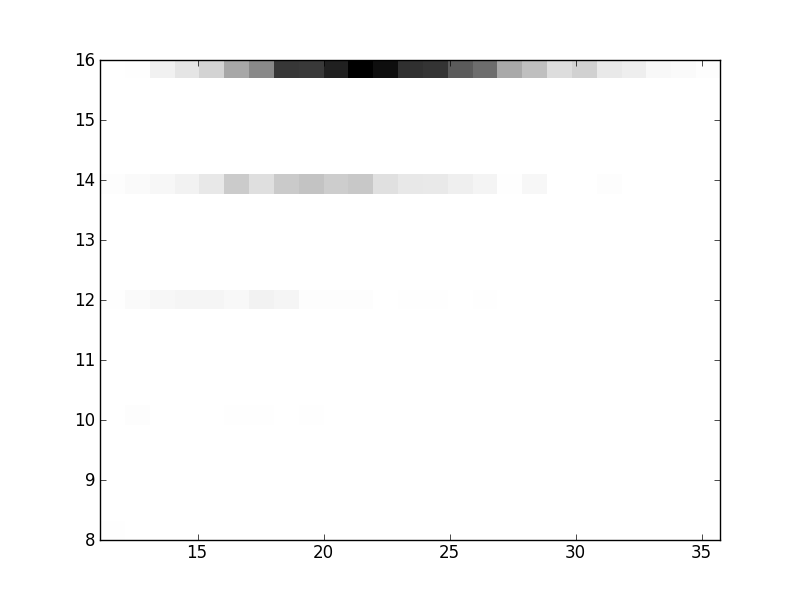 |
| Nunsp p=1 | | | | |  |  |
| Nunsp p=2 | | | | | |  |









