Cophenetic metrics for phylogenetic trees, after Sokal and Rohlf
Correlation between metrics over phylogenetic trees with 100 leaves (Simulation)
Non-necessary binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 | 1.0 | 0.987184213681 | 0.731755302684 | 0.753918430041 | 0.0915564172216 |
| Coph p=2 | | 1.0 | 0.780030404318 | 0.803423126874 | 0.0883895521784 |
| Nsp p=1 | | | 1.0 | 0.990944338279 | 0.132030406364 |
| Nsp p=2 | | | | 1.0 | 0.118336282301 |
| RF | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 |  | 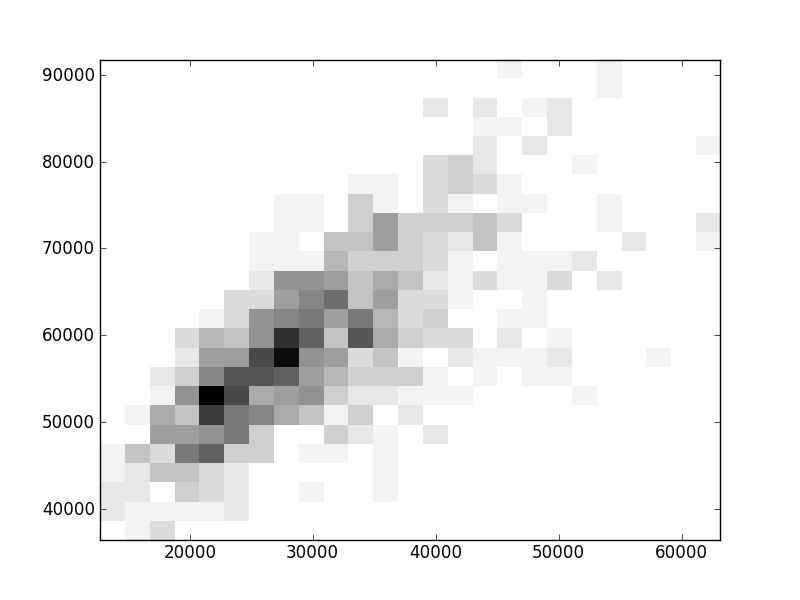 | 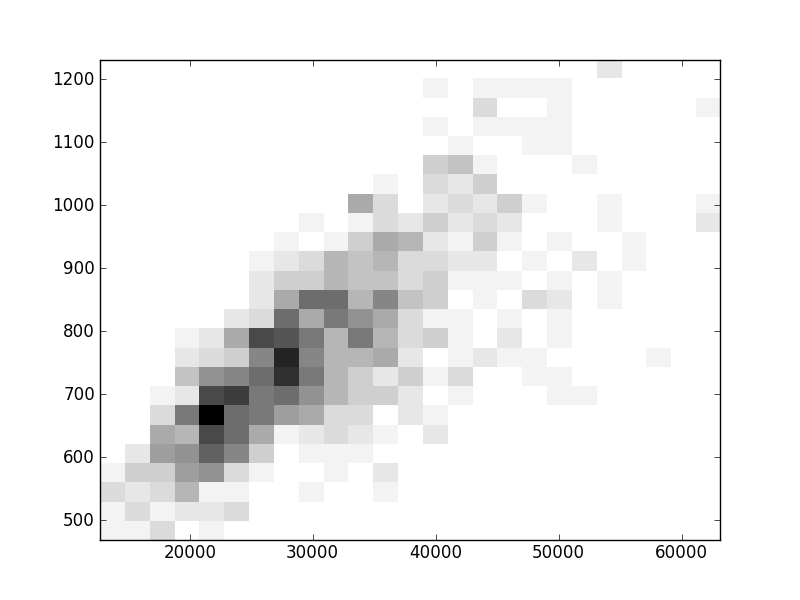 |  |
| Coph p=2 | |  |  |  |
| Nsp p=1 | | | 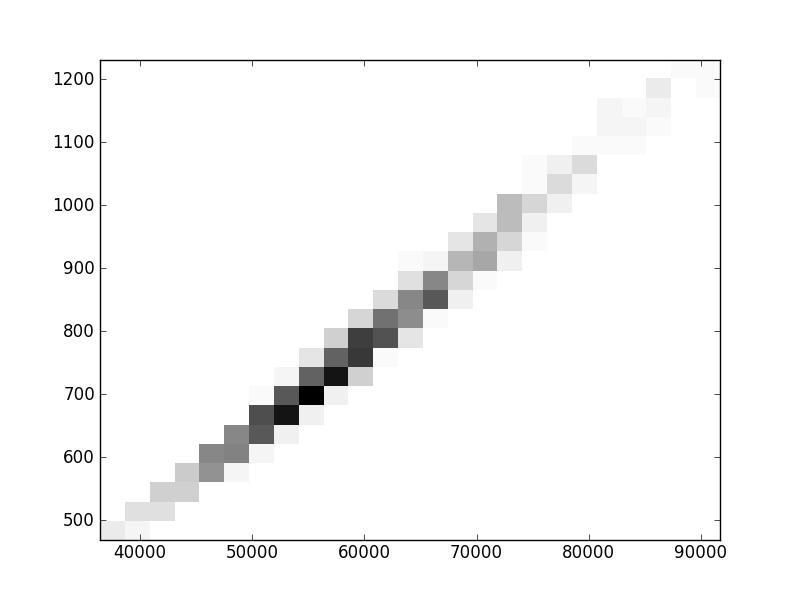 |  |
| Nsp p=2 | | | |  |
Binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 | 1.0 | 0.98693290264 | 0.735760760774 | 0.764001073882 | 0.447139284325 | 0.448265406888 | -0.000800350922793 |
| Coph p=2 | | 1.0 | 0.786239071788 | 0.815571840795 | 0.513306458821 | 0.514362894979 | 0.00328065497692 |
| Nsp p=1 | | | 1.0 | 0.989786590753 | 0.83454385226 | 0.833300531931 | 0.021278127515 |
| Nsp p=2 | | | | 1.0 | 0.804123995169 | 0.805705674073 | 0.0194132768584 |
| Nunsp p=1 | | | | | 1.0 | 0.998478492949 | 0.0126432620494 |
| Nunsp p=2 | | | | | | 1.0 | 0.0123911767917 |
| RF | | | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 |  |  |  | 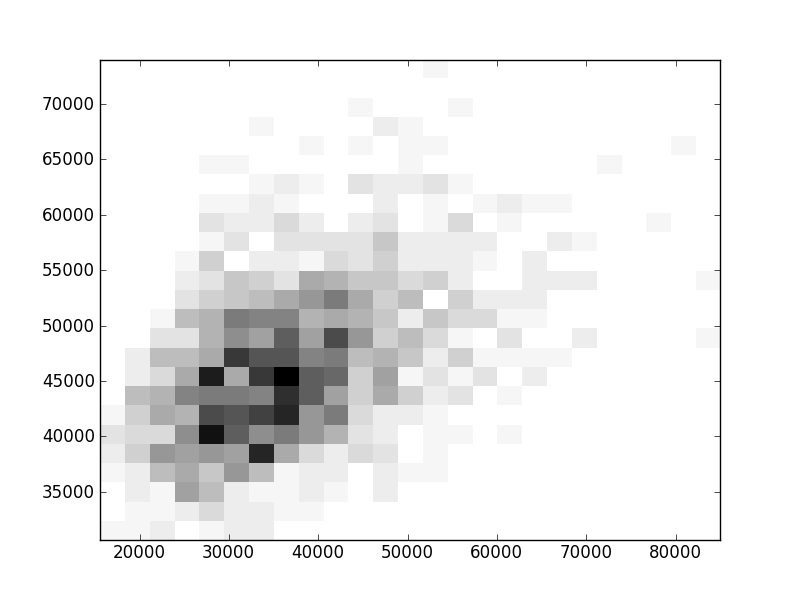 |  |  |
| Coph p=2 | | 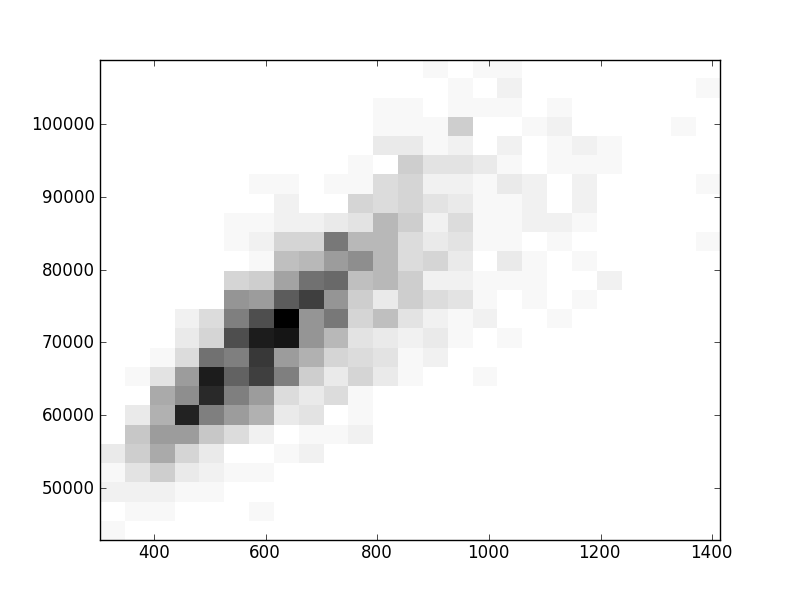 |  |  |  |  |
| Nsp p=1 | | |  |  | 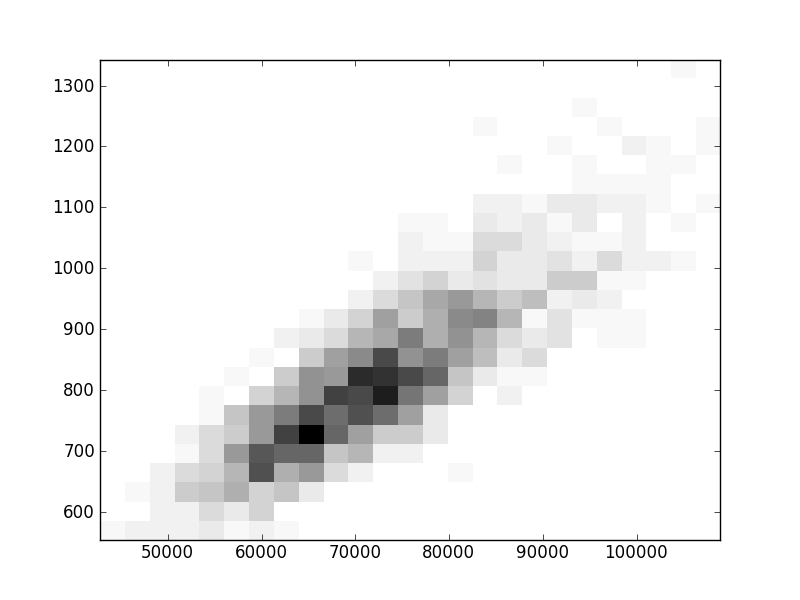 |  |
| Nsp p=2 | | | | 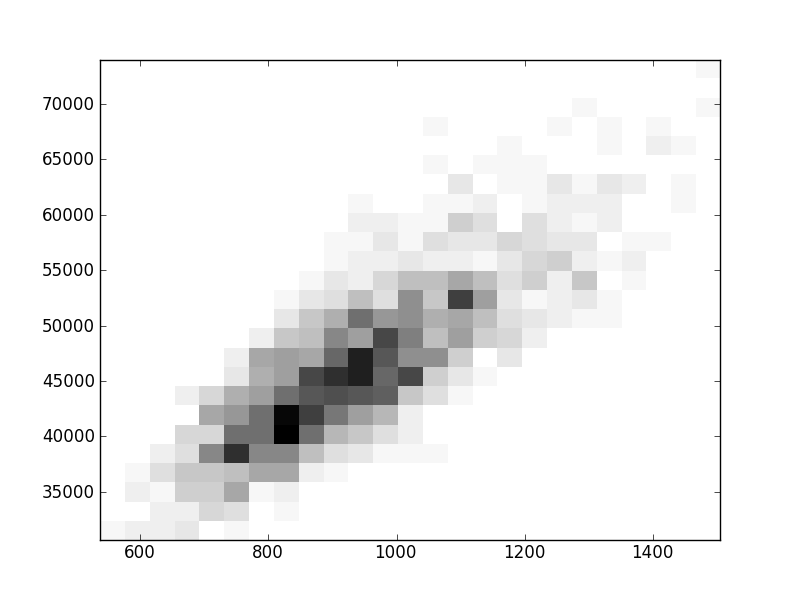 |  |  |
| Nunsp p=1 | | | | |  |  |
| Nunsp p=2 | | | | | |  |









