Cophenetic metrics for phylogenetic trees, after Sokal and Rohlf
Correlation between metrics over phylogenetic trees with 20 leaves (Simulation)
Non-necessary binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 | 1.0 | 0.983363612097 | 0.759259439454 | 0.792567043309 | 0.210551476549 |
| Coph p=2 | | 1.0 | 0.795226483261 | 0.83504824488 | 0.218609581096 |
| Nsp p=1 | | | 1.0 | 0.980020065785 | 0.347661987395 |
| Nsp p=2 | | | | 1.0 | 0.315273968022 |
| RF | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 |  |  |  |  |
| Coph p=2 | | 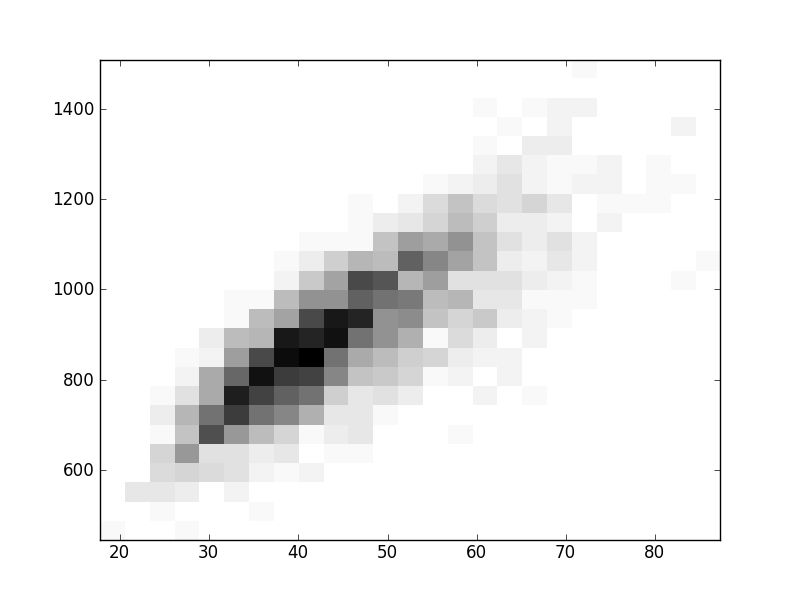 |  |  |
| Nsp p=1 | | |  |  |
| Nsp p=2 | | | | 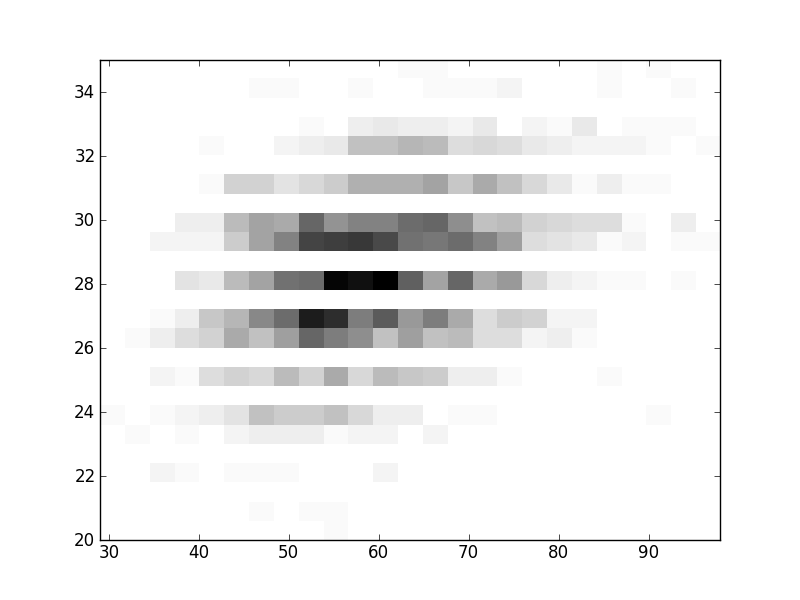 |
Binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 | 1.0 | 0.984801051267 | 0.80391737204 | 0.827696709215 | 0.287948541607 | 0.304452682441 | 0.0594019803224 |
| Coph p=2 | | 1.0 | 0.833753454014 | 0.861866387428 | 0.336461187618 | 0.35478495114 | 0.0536734711025 |
| Nsp p=1 | | | 1.0 | 0.979937742616 | 0.545360041956 | 0.553120178402 | 0.0705055764874 |
| Nsp p=2 | | | | 1.0 | 0.537981985168 | 0.551764078642 | 0.060249553523 |
| Nunsp p=1 | | | | | 1.0 | 0.988363181449 | 0.124553581349 |
| Nunsp p=2 | | | | | | 1.0 | 0.125119269091 |
| RF | | | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 |  | 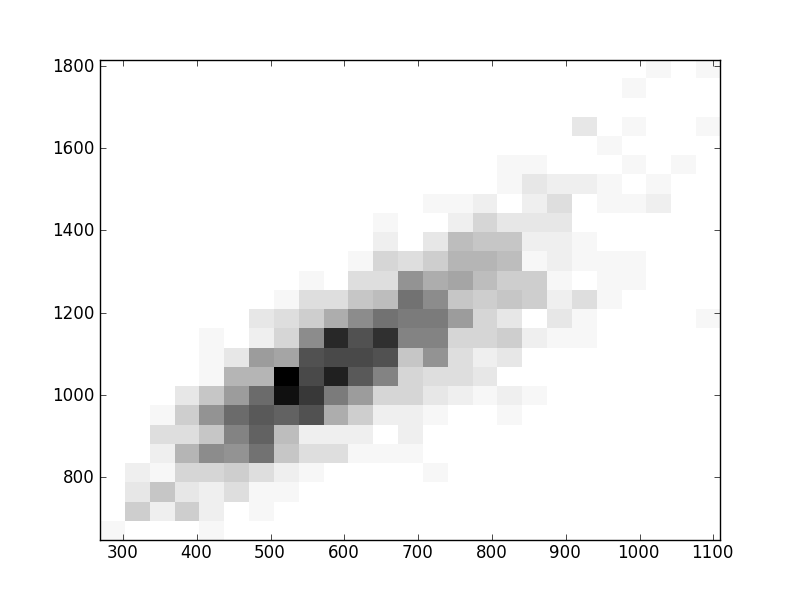 |  | 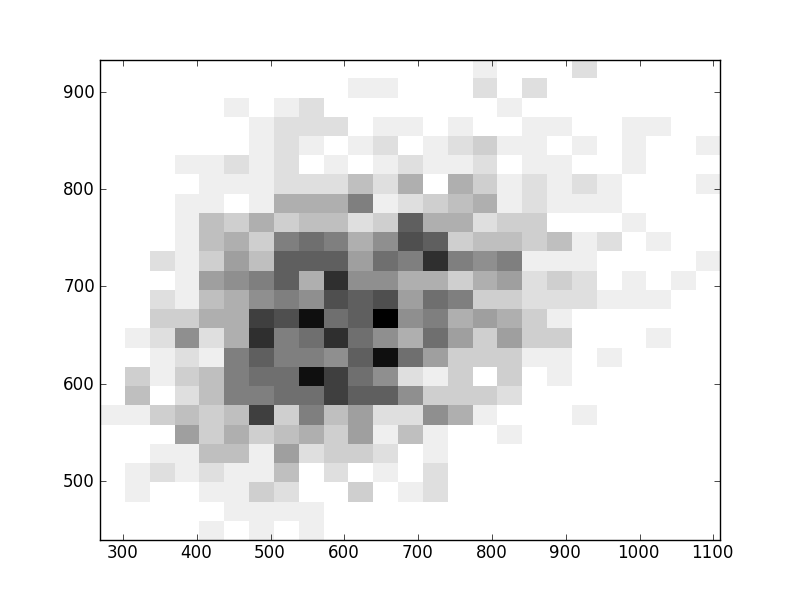 |  |  |
| Coph p=2 | |  | 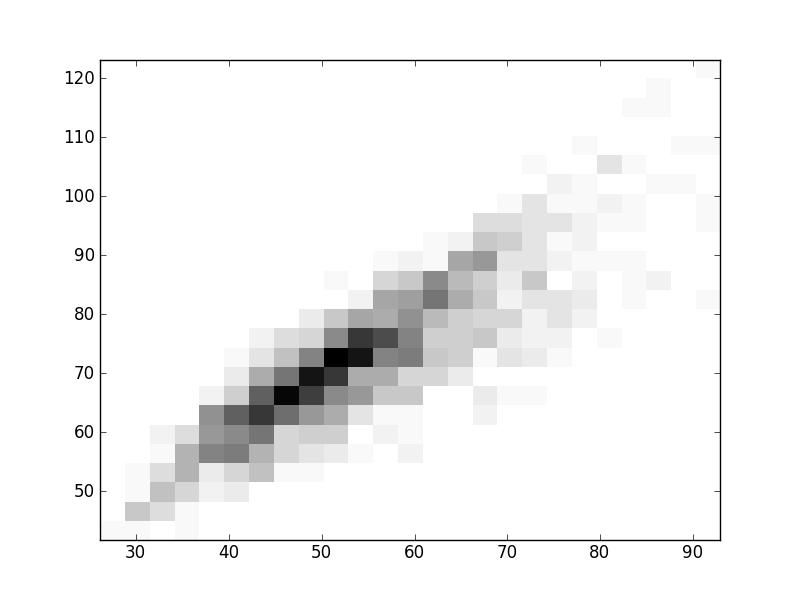 |  | 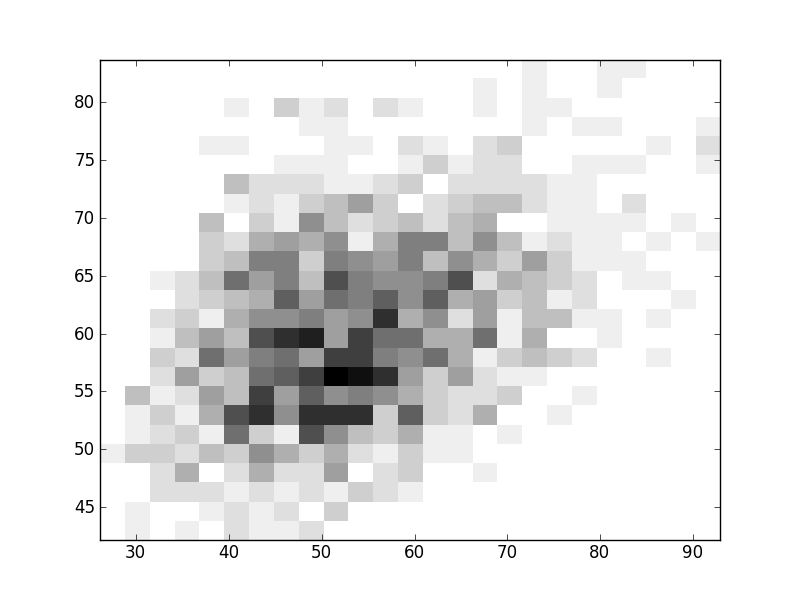 |  |
| Nsp p=1 | | |  |  |  | 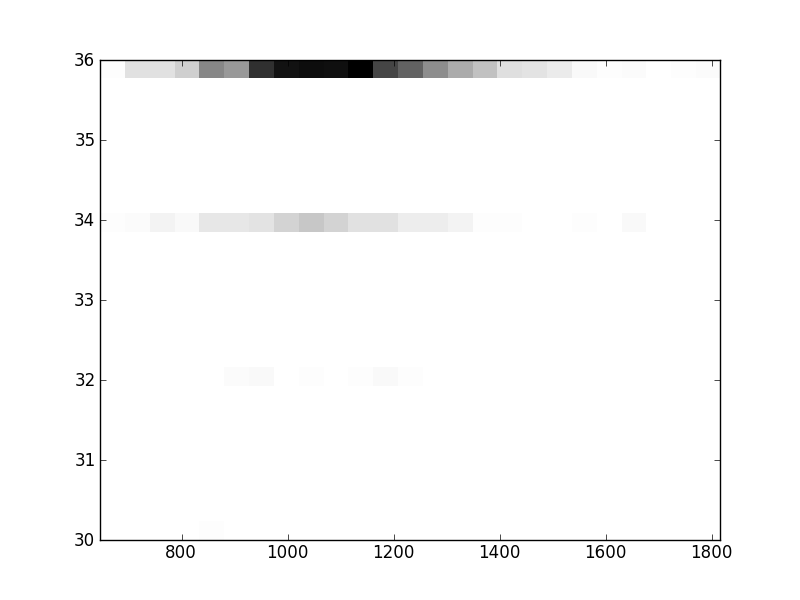 |
| Nsp p=2 | | | |  |  |  |
| Nunsp p=1 | | | | |  |  |
| Nunsp p=2 | | | | | |  |









