Cophenetic metrics for phylogenetic trees, after Sokal and Rohlf
Correlation between metrics over phylogenetic trees with 30 leaves (Simulation)
Non-necessary binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 | 1.0 | 0.984699873098 | 0.764638332165 | 0.789865085425 | 0.192414199228 |
| Coph p=2 | | 1.0 | 0.805949159235 | 0.834840636752 | 0.211302125525 |
| Nsp p=1 | | | 1.0 | 0.983774033454 | 0.316329637799 |
| Nsp p=2 | | | | 1.0 | 0.278542135769 |
| RF | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | RF |
| Coph p=1 |  |  |  | 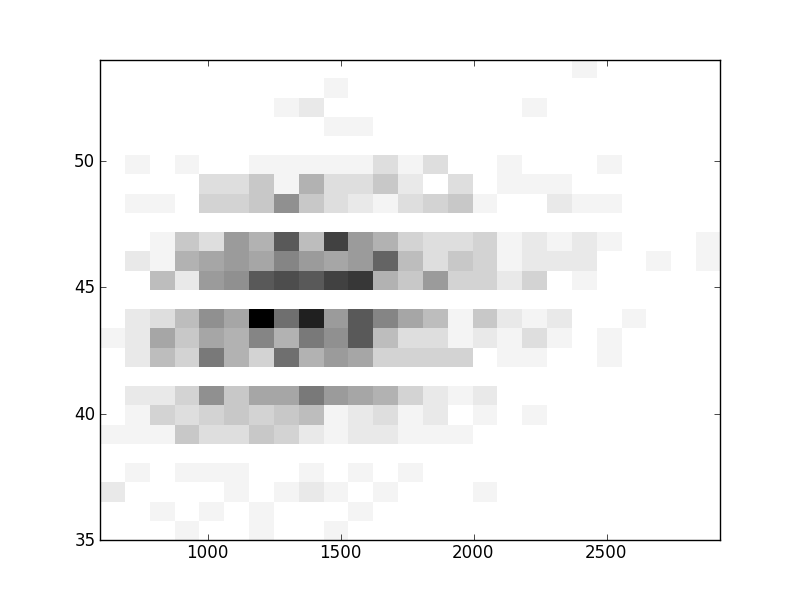 |
| Coph p=2 | |  |  |  |
| Nsp p=1 | | |  |  |
| Nsp p=2 | | | | 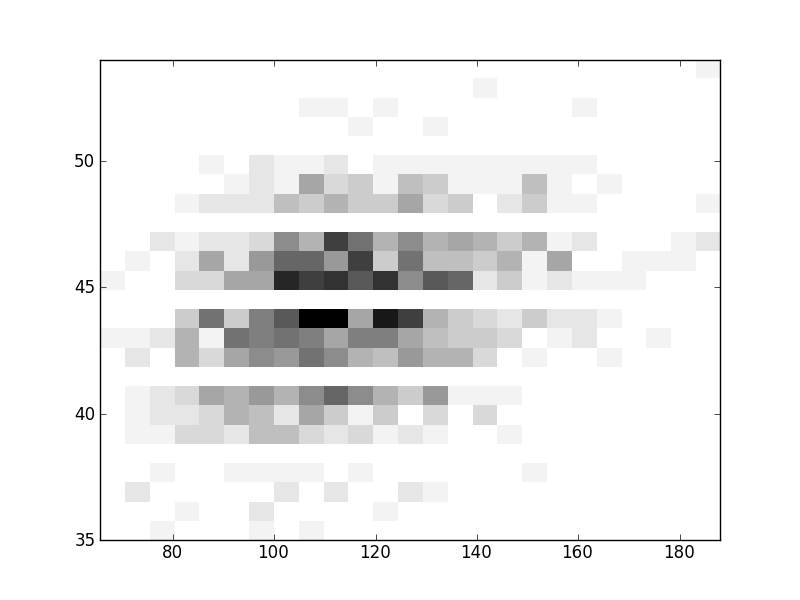 |
Binary phylogenetic trees
Spearman's correlation between cophenetic, nodal and RF metrics.
| | Cophenetic p=1 | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 | 1.0 | 0.986120600676 | 0.783537794638 | 0.806147206471 | 0.358104980459 | 0.359763733722 | 0.0388768726685 |
| Coph p=2 | | 1.0 | 0.815878082571 | 0.843766709703 | 0.420427153216 | 0.423571981371 | 0.0449493853787 |
| Nsp p=1 | | | 1.0 | 0.985469558203 | 0.676811988295 | 0.679564722899 | 0.067682439249 |
| Nsp p=2 | | | | 1.0 | 0.663065721816 | 0.669391148249 | 0.0624325341805 |
| Nunsp p=1 | | | | | 1.0 | 0.994472468785 | 0.114597723131 |
| Nunsp p=2 | | | | | | 1.0 | 0.110089202745 |
| RF | | | | | | | 1.0 |
2D histograms. A darker squeare means that there are more trees with that relationship.
| | Cophenetic p=2 | Nodalsp p=1 | Nodalsp p=2 | Nodalunsp p=1 | Nodalunsp p=2 | RF |
| Coph p=1 |  | 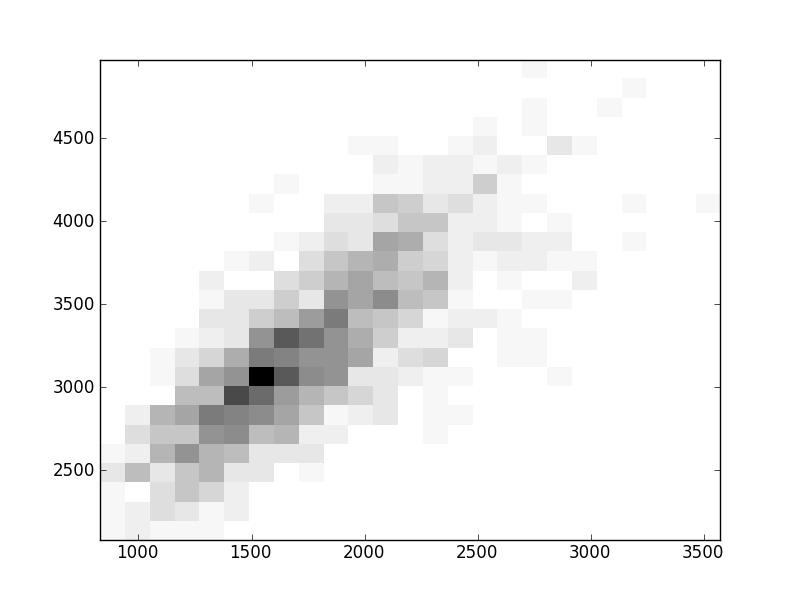 |  |  |  |  |
| Coph p=2 | |  |  |  |  | 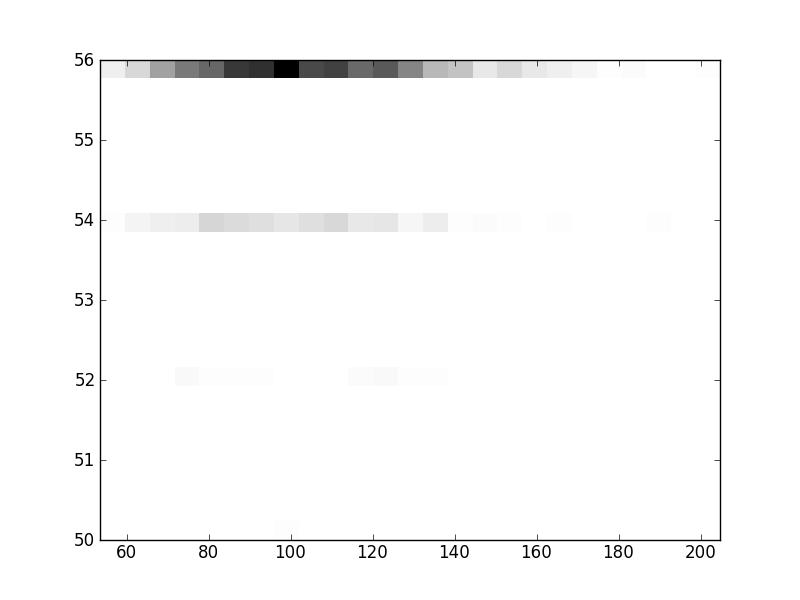 |
| Nsp p=1 | | | 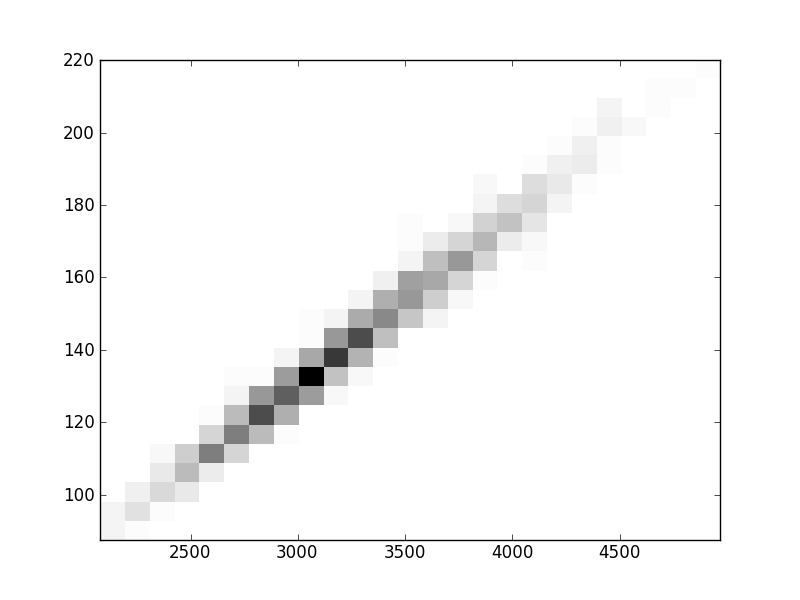 |  |  |  |
| Nsp p=2 | | | | 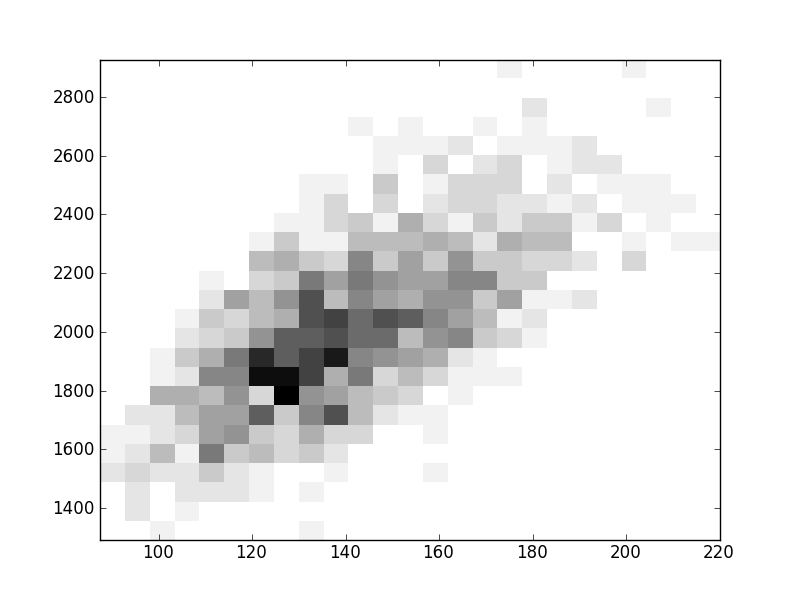 |  |  |
| Nunsp p=1 | | | | |  | 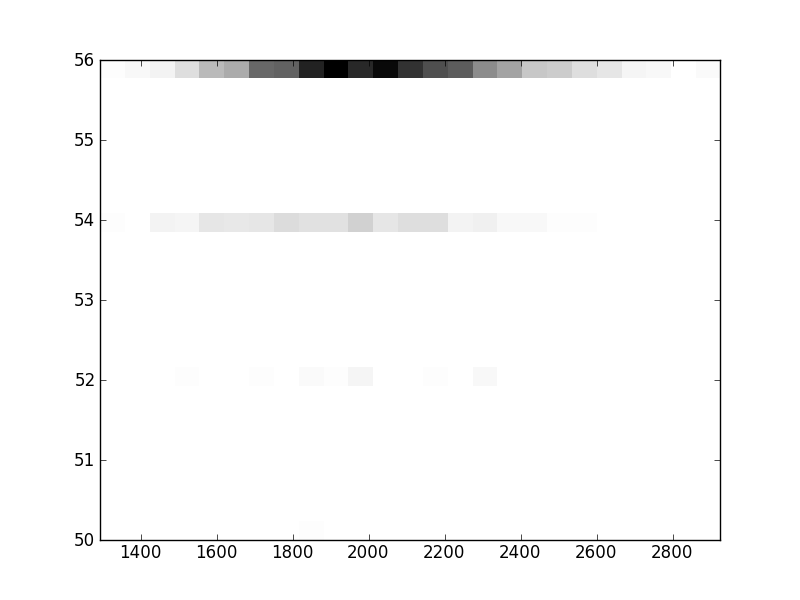 |
| Nunsp p=2 | | | | | |  |









